It is the
time for ancient genomes. A month ago I
read about new ancient samples from England, Hinxton, and saw them to be interesting in terms
of the Finnish history. Those samples are
around 1500-2000 years old, thus being rather suitable for estimating Finnish
western connections. The Finnish
history in Finland is rather short, in the best case bloodlines goes around 2000 years to the past, quite a short
history compared to many southern Europeans.
Data
I use now
the same data I had in my roll-off analyses.
Just to remind you, I made a very strict qualification for Finnish
samples to remove all recent admixtures, meaning the time span from the beginning
of the Swedish era in Finland. All
public western Finnish samples were selected by comparing to my own genealogically
proved samples and outliers were removed.
Software
I used
Reich’s three population test (qp3Pop) with default settings.
Results
Before
going forward some words of caution.
After testing with larger data I realized that also qp3Pop makes an
assumption that less diverse populations are source populations for more
diverse sum populations, in other words diverse populations are usually
composed from several less diverse populations. This is not true and is a rather mechanic
perception. In genetics the process can
be reverse; a more diverse population can turn in to a less diverse one through
genetic drift. This is important because
just the drift is now what we analyze.
Some
general observations. This
above-mentioned problem doesn’t have effect on ancient samples, because they
lived far before us and they can’t violate causality. However the lack of diversity can overestimate
the admixture.
I have also
some results using preset-day source populations and those results can be
problematic. Nevertheless, despite of
the fact that some Finnish samples are from young isolations I
assume that my Eastern Finnish samples represent historically most unmixed
Finnic language speakers in my data, keeping in mind however that I have no
Finnic (Baltic Finnish) speakers from Russia. Additional samples from Russia could give
information about possible admixtures of East Finns.
Abbreviations:
AS -
Hinxton Anglo-Saxons BR - Hinxton Iron Age Briton EF – East Finns WF – West Finns
PL – Poles LT
– Lithuanians EE – Estonians MA – Maris CH – Chuvashes NR – Norwegians
MR – Mordvas BU - Belarussians
Negative
F_3 values mean likely that the target has admixtures of both source
populations.
The Western Finnish map shows high ancient admixtures, especially the Anglo-Saxon - East Finnish admixture among them is outstanding! Estonians show admistures with almost all their neighbors, which can point out continuous migrations to Estonia through the history.
Another way
to find out speculative admixture of source populations is to pick the least probable target population,
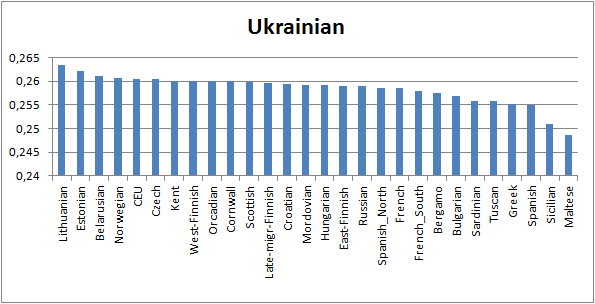
in this case African Pygmies. Using this
method we see that the most Anglo-Saxon-like are Norwegians and the most Iron
Age Briton-like are Lithuanians. West Finns
are the third on the Anglo-Saxon axis. This
probably means that West Finns have Anglo-Saxon-like ancestry, or Anglo Saxons
had common Fennoscandinavian ancestry with West Finns. All those owning more Iron Age British ancestry
than West Finns (NR, LT, EE, PL) likely have more ancient Celtic ancestry from
Central Europe.

edit Su 16.11.2014
I thought that it would be interesting to know more about the western outlier group, the Finns who are more western on PCA plots than the genealogically proved West Finnish group. This is done by comparing both western Finnish groups to East Finns, that is to say the East Finns are a fixed landmark on which to base the comparison. The result shows mixed results with negative f3-stat F3(WF;EF,test) and f3(WF2;EF,test) where "test" includes Estonians, Swedes, Iron Age Britons and ancient Anglo-Saxons.
The result shows that western outliers show more Iron Age Briton, more Estonian and more Swedish ancestry than the genealogical western group, but they show little less ancient Anglo-Saxon ancestry than the genealogical western group. The result also proves that both western Finnish groups have significant Eastern Finnish -like ancestry.
The abbreviation "SE" stands for three Swedish samples who show only very little Finnish ancestry at 23andme's Ancestry Composition.
edit. Mon 17.11.2014
Another graphic showing the Swedish - ancient Anlo-Saxon ratio among Finnish individuals, both admixture gotten by 3Pop-software. The East Finnish group is used as a fixed landmark. The individual difference between AS and SE was used as a sort key and the trend line shows linear difference. I would have done also comparisons to other populations, like to Estonians, but the difference in SNP-sets made an individual level comparison impossible.
Although ancient Anglo-Saxon and Swedish admixtures follow each other, my judgement is that the bigger the AS is compared to the SE, the bigger is the ancient admixture, and vice vesa.