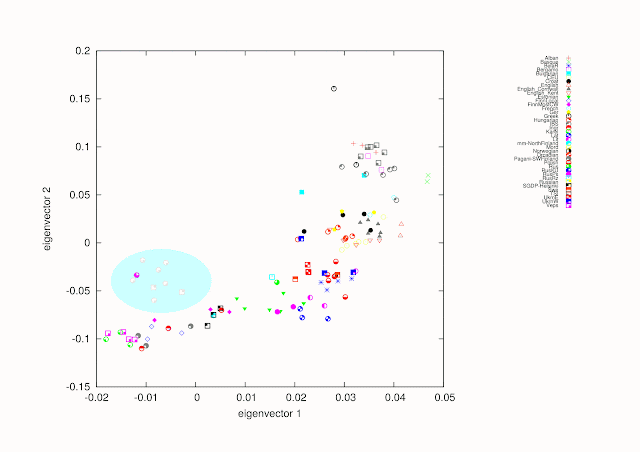
Here, as a showcase of the new data two PCA prints and some comments. Instead of making a central continental European picture I included four outgroups to see the effect. Those outgroups are Armenians, Mongolians, Sardinians and Saamis. For the present individual names are picked straight from original sources and can be somewhat ambiguous.
As we can see we have several clusters, which makes possible to evaluate the data. For example Scandinavians of SGDP and Pagani et al. cluster with East Europeans.
Personally I don't give much attention to PCA-figures, because the result depends on the selected samples, amounts, ratios between populations sizes, about how mixed are individuals etc. My upcoming high resolution tests will be much more interesting.
Added time 12:50
If someone is interested in how Mordvas locate on this map. They are very similar to North Russians and RusKU (Pagani et al.) and move towards Mongolians. Baltic Finns moves towards Saamis. Sorry, GIMP makes something unwanted with colors.
I